Genome-wide Identification and Expression Analysis of AaGRF Gene Family in Artemisia argyi
-
摘要:
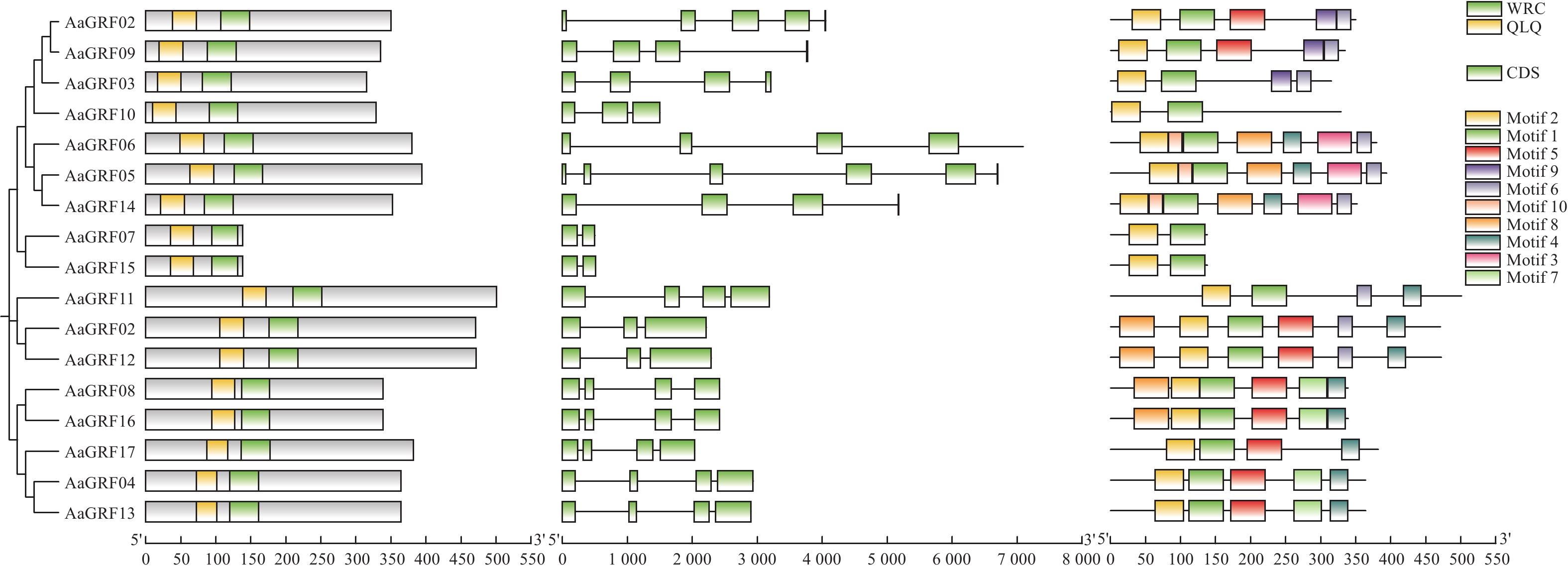
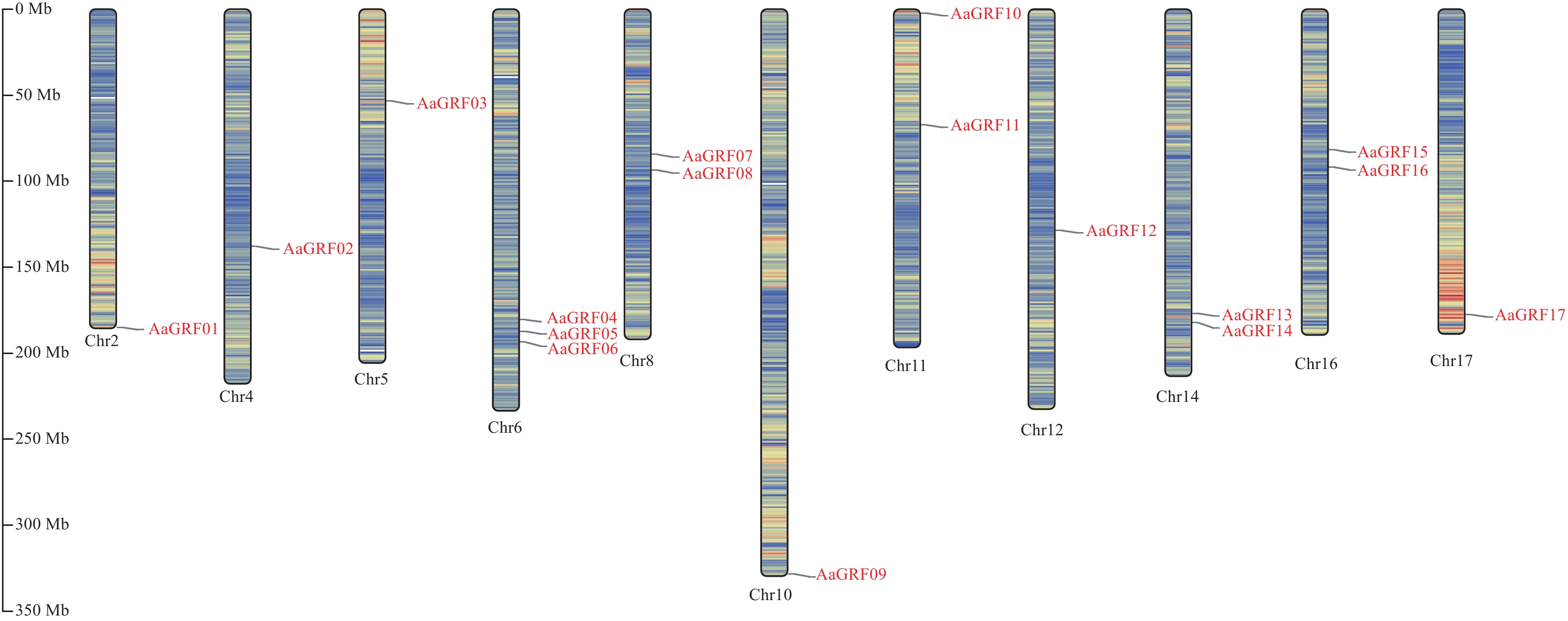
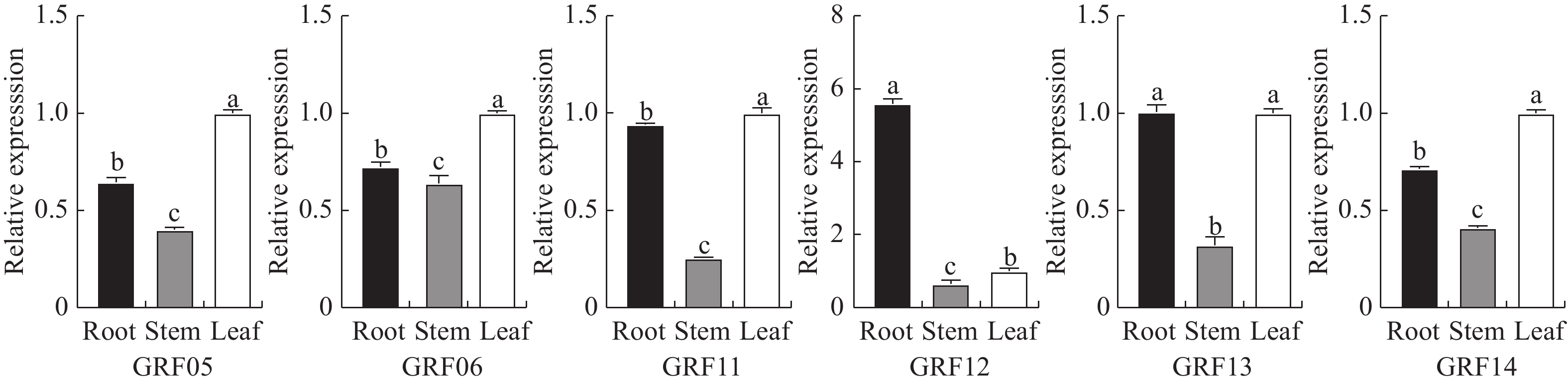
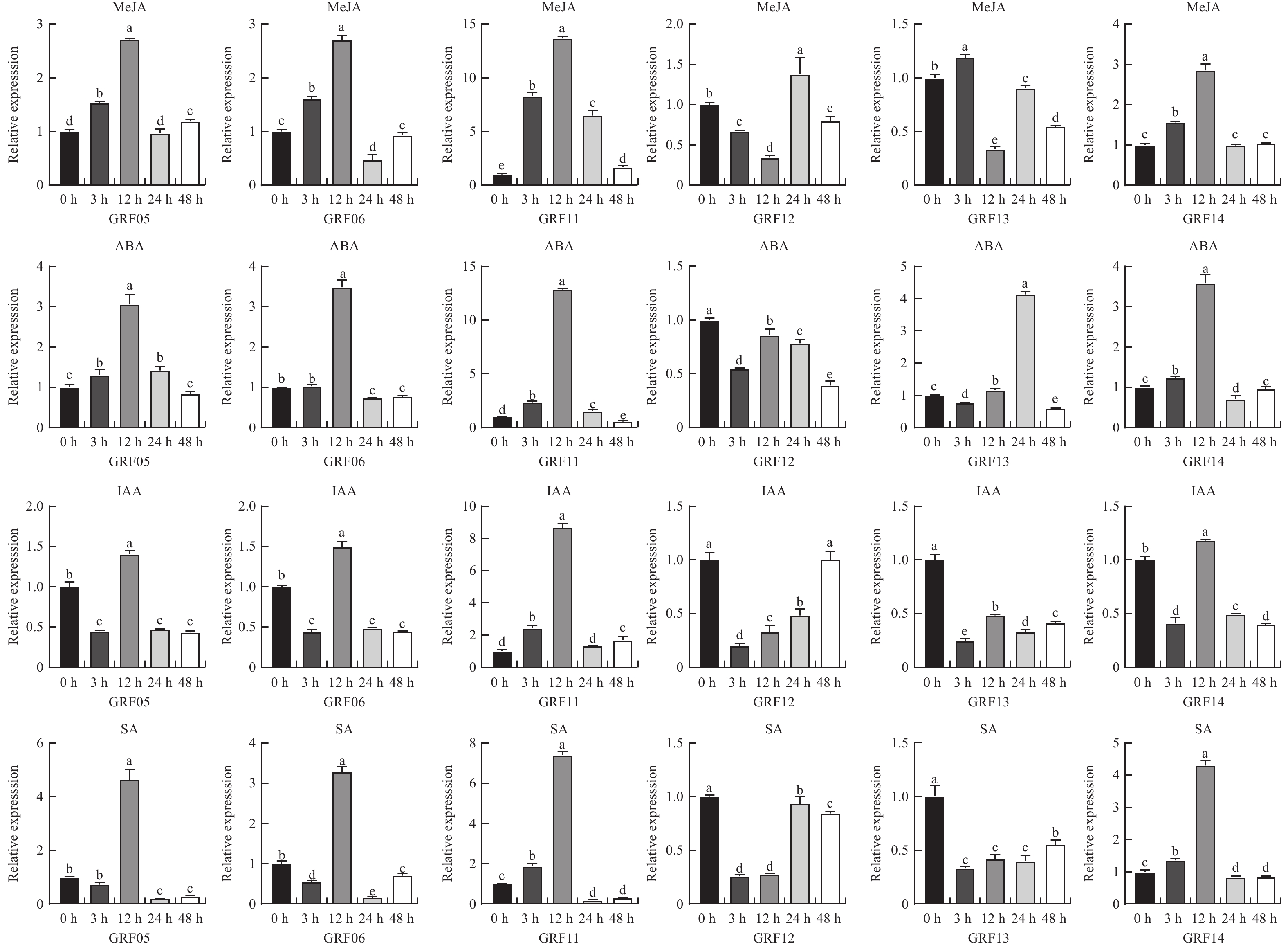
目的 生长调节因子(Growth-regulating factors, GRF)是植物的一种特有蛋白,深入了解艾GRF基因家族的生物学特性,对艾的生长发育和逆境胁迫有着重要作用。 方法 本研究对艾GRF基因家族进行理化性质、进化、基因结构以及表达等进行生物信息学进行分析,进而通过转录组数据和实时荧光定量PCR(qRT-PCR)技术分析基因家族各成员在不同组织、逆境胁迫以及响应不同激素的表达模式进行分析。 结果 从艾基因组中共鉴定出17个GRF家族成员,理化鉴别显示均为亲水蛋白;系统发育进化树分为四个亚家族;motif分析表明进化树中同一分支的基因具有相似或相同的保守基序;AaGRF基因家族成员随机分布在11条不同的染色体上;AaGRF基因家族启动子含有多种逆境和激素响应元件;AaGRF基因家族在响应盐碱胁迫时表现出不同的表达趋势;该家族成员在艾的根、茎、叶中的表达模式具有组织特异性;经脱落酸、吲哚乙酸、水杨酸和茉莉酸甲酯处理后,AaGRF05、AaGRF06、AaGRF11和AaGRF14在处理后的12h基因表达量最高,这表明大部分基因在前期参与不同激素的调控。 结论 艾GRF基因家族参与艾生长发育、激素与逆境胁迫调节,为下一步全面探究GRF家族功能提供一定的理论依据。 Abstract:Objective Growth-regulating factors (GRF) is a unique protein of plants, which plays an important role in mugwort growth and stress. It plays an important role in Artemisia argyi growth and stress stress. Method To understand the biological characteristics of the GRF gene family in Artemisia argyi, this study conducted bioinformatic analysis of physical and chemical properties, gene structure, evolution and expression of the GRF gene family; analyzed the expression patterns of the members of the gene family in different tissues, stress and different hormones using transcriptomic data and real-time PCR (qRT-PCR). Result A total of 17 GRF family members were identified from the A. argyi genome, physicochemical identification showed that all are hydrophilic proteins; The phylogenetic evolutionary tree falls into four subfamilies; The motif analysis indicated that genes of the same clade in the evolutionary tree have a similar or the same conserved motif; The AaGRF gene family members are randomly distributed on 11 different chromosomes; The AaGRF gene family promoter contains multiple stress and hormone-responsive elements; The AaGRF gene family showed different trends in response to salinity stress; The expression pattern of this family members in the roots, stems and leaves of A. argyi is tissue-specific; After treatment with abscisic acid, indoacetic acid, salicylic acid and methyl jasmonate, AaGRF05, AaGRF06, AaGRF11 and AaGRF14 showed the highest gene expression at 12 hour after treatment, indicating that most of the genes are involved in regulation during prophase. Conclusion In conclusion, this paper shows that the GRF gene family participates in the regulation of A. argyi growth and stresses, which provides a certain theoretical basis for the comprehensive exploration of the function of GRF family in the next step research. -
Key words:
- Artemisia argyi /
- GRF gene family /
- expression analysis /
- hormone respone
-
表 1 AaGRF基因家族蛋白理化性质
Table 1. Physiochemical properties of the AaGRF gene family proteins
基因名称
Gene name基因组ID
genome ID长度
length/aa分子量
molecular weight /Da等电点
Isoelectric point(pl)不稳定指数
Instability index脂肪族指数
Aliphatic index疏水指数
Hydrophobic indexAaGRF01 Aarg02G031430.1 350 39886.41 8.35 51.62 50.43 −0.921 AaGRF02 Aarg04G019450.1 471 50852.5 9.07 41.98 64.86 −0.547 AaGRF03 Aarg05G013120.1 315 36666.98 7.26 71.34 58.83 −0.9 AaGRF04 Aarg06G029080.1 364 40414.66 8.87 48.27 52.77 −0.91 AaGRF05 Aarg06G030040.1 394 43558.38 9 48.32 57.92 −0.717 AaGRF06 Aarg06G030730.1 380 42917.36 8.13 56.92 54.92 −0.874 AaGRF07 Aarg08G013810.1 138 15560.65 7.66 52.91 65.72 −0.541 AaGRF08 Aarg08G014980.1 339 38274.79 8.02 49.95 43.42 −0.871 AaGRF09 Aarg10G061780.1 335 38176.52 8.79 49.88 52.39 −0.87 AaGRF10 Aarg11G000710.1 329 37157.12 8.97 47 52.52 −0.829 AaGRF11 Aarg11G015240.1 501 54688.51 7.24 42.86 59.38 −0.635 AaGRF12 Aarg12G018140.1 472 51001.69 9.16 42.64 64.51 −0.56 AaGRF13 Aarg14G027820.1 364 40379.65 8.72 45.86 51.43 −0.898 AaGRF14 Aarg14G028850.1 352 38751.75 8.44 51.96 54.29 −0.709 AaGRF15 Aarg16G013730.1 138 15512.59 7.66 52.9 69.28 −0.509 AaGRF16 Aarg16G015010.1 339 38280.78 7.66 49.46 45.13 −0.853 AaGRF17 Aarg17G035750.1 382 41725.01 6.67 45.33 56.18 −0.678 -
[1] 简梨娜, 宋学丽, 郭江涛, 等. 艾草的化学成分及临床应用 [J]. 化学工程师, 2021, 35(7):58−62.JIAN L N, SONG X L, GUO J T, et al. Chemical composition and clinical application of Artemisia argyi [J]. Chemical Engineer, 2021, 35(7): 58−62. (in Chinese) [2] 李晓红. 蕲艾的采集储藏及药用价值的探讨 [J]. 中国医药指南, 2012, 10(27):615−616.LI X H. Discussion on the collection, storage and medicinal value of Artemisia argyi [J]. Guide of China Medicine, 2012, 10(27): 615−616. (in Chinese) [3] 马超, 原佳乐, 张苏, 等. GRF转录因子对植物生长发育及胁迫响应调控的分子机制 [J]. 核农学报, 2017, 31(11):2145−2153. doi: 10.11869/j.issn.100-8551.2017.11.2145MA C, YUAN J L, ZHANG S, et al. The molecular mechanisms of growth-regulating factors (GRFs) in plant growth, development and stress response [J]. Journal of Nuclear Agricultural Sciences, 2017, 31(11): 2145−2153. (in Chinese) doi: 10.11869/j.issn.100-8551.2017.11.2145 [4] 邱玮茜, 王帅, 鞠青, 等. 鳗草GRF基因家族的鉴定与生物信息学分析 [J]. 浙江海洋大学学报(自然科学版), 2022, (1):31−37.QIU W Q, WANG S, JU Q, et al. Genome-wide identification and bioinformatics analysis of growth-regulating factor gamily in Zostera marina [J]. Journal of Zhejiang Ocean University (Natural Science), 2022(1): 31−37. (in Chinese) [5] CHOI D, KIM J H, KENDE H. Whole genome analysis of the OsGRF gene family encoding plant-specific putative transcription activators in rice (Oryza sativa L. ) [J]. Plant and Cell Physiology, 2004, 45(7): 897−904. doi: 10.1093/pcp/pch098 [6] . 张宴萍, ?, ?等, 马铃薯GRF家族的全基因组鉴定与生物信息学分析[J]. 分子植物育种, 2024(?): 1-9.ZHANG Y P , ?, ?et al. , genome-wide identification and bioinformatics analysis of the GRF gene family in potato [J]. Molecular plant breeding, 2024(?): 1-9. (in Chinese) [7] KHATUN K, ROBIN A H K, PARK J I, et al. Molecular characterization and expression profiling of tomato GRF transcription factor family genes in response to abiotic stresses and phytohormones [J]. International Journal of Molecular Sciences, 2017, 18(5): 1056. doi: 10.3390/ijms18051056 [8] WANG P, XIAO Y, YAN M, et al. Whole-genome identification and expression profiling of growth-regulating factor (GRF) and GRF-interacting factor (GIF) gene families in Panax ginseng [J]. BMC Genomics, 2023, 24: 334. doi: 10.1186/s12864-023-09435-w [9] KUIJT S J H, GRECO R, AGALOU A, et al. Interaction between theGROWTH-REGULATING FACTOR and KNOTTED1-LIKE HOMEOBOX Families of transcription factors [J]. Plant Physiology, 2014, 164(4): 1952−1966. doi: 10.1104/pp.113.222836 [10] LI S C, GAO F Y, XIE K L, et al. The OsmiR396c-OsGRF4-OsGIF1 regulatory module determines grain size and yield in rice [J]. Plant Biotechnology Journal, 2016, 14(11): 2134−2146. doi: 10.1111/pbi.12569 [11] LIAN C L, LAN J X, MA R, et al. Genome-wide analysis of aux/IAA gene family in Artemisia argyi: Identification, phylogenetic analysis, and determination of response to various phytohormones [J]. Plants, 2024, 13(5): 564. doi: 10.3390/plants13050564 [12] 易小哲, 邬兰, 向丽, 等. 艾Artemisia argyi实时荧光定量PCR内参基因筛选 [J]. 中国中药杂志, 2022, 8(3):659−667.YI X Z, WU L, XIANG L, et al. Screening of reference genes for quantitative real-time PCR in Artemisia argyi [J]. China Journal of Chinese Materia Medica, 2022, 8(3): 659−667. (in Chinese) [13] 薛正刚, 王树杰, 杨永乾, 等. 大麦GRF家族的基因组鉴定及生物信息学分析 [J]. 分子植物育种, 2021, 19(6):1750−1757.XUE Z G, WANG S J, YANG Y Q, et al. Genome-wide identification and bioinformatics analysis of growth regulating factor(GRF)family in barley [J]. Molecular Plant Breeding, 2021, 19(6): 1750−1757. (in Chinese) [14] 陈淑颖, 李一凡, 曾伟伟, 等. 圆齿野鸦椿YABBY基因家族鉴定及功能分析 [J]. 森林与环境学报, 2023, 43(3):303−310.CHEN S Y, LI Y F, ZENG W W, et al. Identification and functional analysis of the YABBY gene family in Euscaphis konishii [J]. Journal of Forest and Environment, 2023, 43(3): 303−310. (in Chinese) [15] SINGH K. Transcription factors in plant defense and stress responses [J]. Current Opinion in Plant Biology, 2002, 5(5): 430−436. doi: 10.1016/S1369-5266(02)00289-3 [16] FENG K, HOU X L, XING G M, et al. Advances in AP2/ERF super-family transcription factors in plant [J]. Critical Reviews in Biotechnology, 2020, 40(6): 750−776. doi: 10.1080/07388551.2020.1768509 [17] DU W X, YANG J F, LI Q, et al. Genome-wide identification and characterization of growth regulatory factor family genes in Medicago [J]. International Journal of Molecular Sciences, 2022, 23(13): 6905. doi: 10.3390/ijms23136905 [18] WANG F D, QIU N W, DING Q, et al. Genome-wide identification and analysis of the growth-regulating factor family in Chinese cabbage (Brassica rapa L. ssp. pekinensis) [J]. BMC Genomics, 2014, 15(1): 807. doi: 10.1186/1471-2164-15-807 [19] KIM J H, CHOI D, KENDE H. The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis [J]. The Plant Journal, 2003, 36(1): 94−104. doi: 10.1046/j.1365-313X.2003.01862.x [20] NOON J B, HEWEZI T, BAUM T J. Homeostasis in the soybean miRNA396–GRFnetwork is essential for productive soybean cyst nematode infections [J]. Journal of Experimental Botany, 2019, 70(5): 1653−1668. doi: 10.1093/jxb/erz022 [21] 张立全, 张浩林, 李丛丛, 等. 谷子GRF基因家族鉴定与分析 [J]. 西南农业学报, 2021, 34(11):2340−2347.ZHANG L Q, ZHANG H L, LI C C, et al. Genome-wide analysis and identification of GRF gene family in foxtail millet (Setaria italica) [J]. Southwest China Journal of Agricultural Sciences, 2021, 34(11): 2340−2347. (in Chinese) [22] 时丕彪, 何冰, 费月跃, 等. 藜麦GRF转录因子家族的鉴定及表达分析 [J]. 作物学报, 2019, 45(12):1841−1850.SHI P B, HE B, FEI Y Y, et al. Identification and expression analysis of GRF transcription factor family of Chenopodium quinoa [J]. Acta Agronomica Sinica, 2019, 45(12): 1841−1850. (in Chinese) [23] . 谷海静等, 陆地棉RVT_3基因家族的全基因组分析[J]. 南通大学学报(自然科学版), 2024: 1-15.GU H J et al. , Genome-wide analysis of RVT _ 3 gene family in Upland cotton[J]. Journal of Nantong University (Natural Science Edition), 2024: 1-15. (in Chinese) [24] HUANG W D, HE Y Q, YANG L, et al. Genome-wide analysis of growth-regulating factors (GRFs) in Triticum aestivum [J]. PeerJ, 2021, 9: e10701. doi: 10.7717/peerj.10701 -







 下载:
下载: